|
īecause of this, new influenza vaccines must be formulated on a biannual basis for upcoming flu seasons in the Northern and Southern hemispheres. Indeed, the most devastating human influenza epidemics, including the 1918 Spanish Flu, the 1968 Hong Kong flu, the 2004 Bird Flu and the 2009 Swine Flu, were all the result of human/non-human influenza A strains that had recombined. While all influenza viruses undergo antigenic drift, only the influenza A viruses are able to undergo antigenic shift due to their extensive animal reservoirs, which provide the viral diversity to facilitate such events. By shifting the epitopes of their surface antigens so drastically, influenza A strains are able to catch the immune system completely off-guard and rapidly proliferate through human populations. This can result in highly advantageous gain-of-function mutations that confer the virus with new disease characteristics or host tropisms. On the other hand, antigenic shift is characterised by abrupt changes to HA and NA through genome reassortment with other influenza subtypes that occurs when two different influenza viruses co-infect the same cell. This allows HA’s epitopes to quickly change and escape host recognition, without hindering influenza’s ability to gain entry to cells. 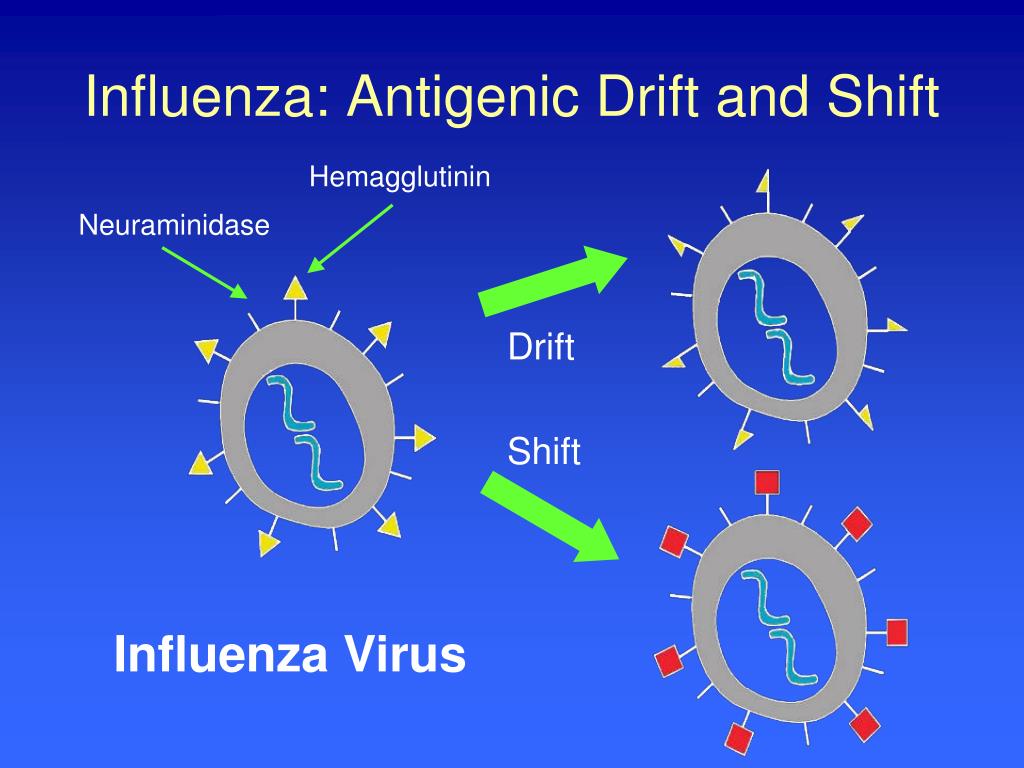
As HA contains various structurally tolerant regions and influenza RNA polymerase lacks proofreading activity, HA mutations can quickly accumulate. Influenza has achieved this to great effect through a potent combination of antigenic drift and shift: antigenic drift is a continuous process in which the development of adaptive immunity by hosts creates selection pressures for mutations in influenza’s Haemagglutinin proteins. This can largely be attributed to their host diversity and ability to mutate. Beyond being RNA viruses with an innate ability to quickly generate genetic variants, influenza viruses are especially adept at undergoing significant genomic changes to evade recognition by adaptive immune responses. The influenza viruses are arguably one of the most successful endemic and pandemic human pathogens. Combined, seasonal influenza is thought to cause an estimated one billion infections and 290,000-650,000 deaths, worldwide each year. However, even a run-of-the-mill flu can be severe in otherwise healthy people, and lethal to those who are elderly or frail, with complications including pneumonia, organ failure and secondary infections. 
Influenza infections usually affect the upper respiratory tract, with mild symptoms, including fever, headache and malaise. 
Influenza virus infections (known as the flu) occur in distinct outbreaks of varying severity every year, with periods of peak incidence (known as flu seasons) oscillating between the northern and southern hemispheres on a seasonal basis. For influenza type A, at least 16 highly variable HAs (H1-18) and 9 distinct NAs (N1-11) have been recognised so far the influenza B virus has a similar viral structure to type A, but lacks H/N subtypes. The Orthomyxoviridae family comprises four major types of influenza virus (A–D), which are classified by their NA and M1 proteins the most clinically relevant are the co-circulating influenza A and B viruses. Influenza’s genome comprises eight single-stranded RNAs encased in a protein capsid, surrounded by a lipid bilayer from which the Haemagglutinin (HA), Neuraminidase (NA) and Matrix (M1, M2) proteins protrude. The influenza viruses are single-stranded RNA viruses belonging to the Orthomyxoviridae family, typically 80-120nm in diameter and spherical or filamentous in shape.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
